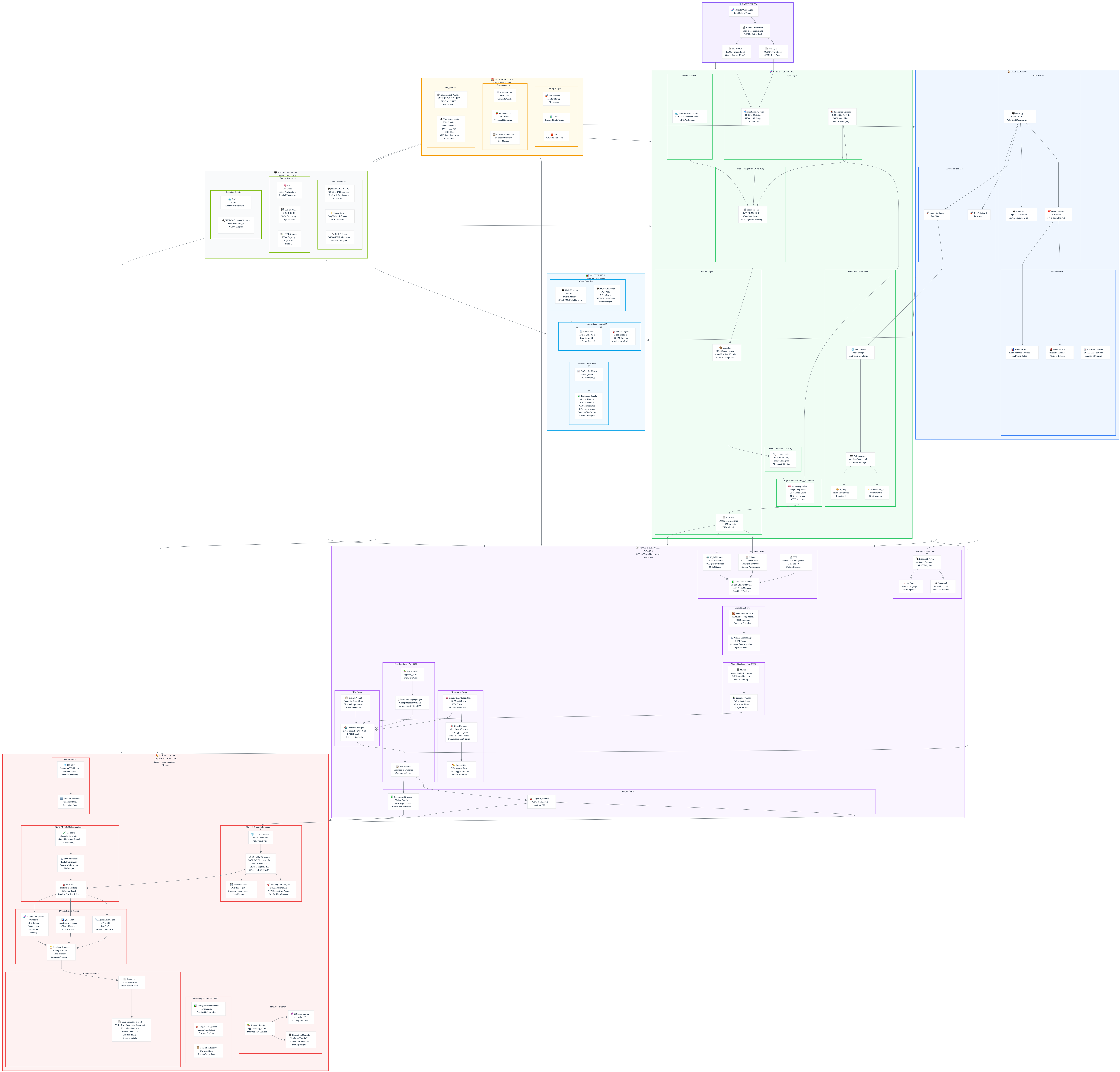
Stage 1: GPU Genomics¶
From raw sequencing data to millions of variants
120 – 240 minutes on DGX Spark
Prerequisite: Stage 0 — Data Acquisition
Before running Stage 1, all required data must be downloaded via setup-data.sh. This includes HG002 FASTQ files (~200 GB), the GRCh38 reference genome, and BWA-MEM2 index. Stage 0 is a one-time step.
What This Stage Does¶
When a patient's DNA is sequenced, the machine produces raw data — billions of short DNA fragments stored in FASTQ files, typically around 200 GB per patient.
Stage 1 transforms this raw data into actionable genetic information:
-
Alignment — Each DNA fragment is mapped back to its position on the human reference genome (like assembling a 3-billion-piece puzzle)
-
Variant Calling — The pipeline identifies where this patient's DNA differs from the reference — these differences are called variants
-
Quality Filtering — AI-powered models (DeepVariant) distinguish real variants from sequencing errors with >99% accuracy
By the Numbers¶
| Metric | Value |
|---|---|
| Input size | ~200 GB FASTQ |
| Reads aligned | 800M – 1.2B |
| Variants discovered | 11.7 million |
| High-quality variants | 3.56 million |
| Accuracy | >99% (DeepVariant) |
| Runtime | 120 – 240 minutes |
The Speed Advantage¶
| Step | Traditional (CPU) | HCLS AI Factory (GPU) |
|---|---|---|
| Alignment (BWA-MEM2) | 12 – 24 hours | 1 – 2 hours |
| Variant Calling | 8 – 12 hours | 1 – 2 hours |
| Total | 1 – 2 days | 2 – 4 hours |
GPU acceleration via NVIDIA Parabricks delivers 10–50x speedup over traditional CPU pipelines.
Technology Stack¶
How It Works¶
flowchart LR
FASTQ["FASTQ\n200 GB raw reads"]
ALIGN["BWA-MEM2\nGPU Alignment"]
DV["DeepVariant\nAI Variant Calling"]
VCF["VCF\n11.7M variants"]
GPU["⚡ NVIDIA GPU"]
FASTQ --> ALIGN --> DV --> VCF
GPU -.->|accelerates| ALIGN
GPU -.->|accelerates| DV
style FASTQ fill:#B3E5FC,stroke:#0288D1
style ALIGN fill:#76B900,stroke:#5a8f00,color:#fff
style DV fill:#76B900,stroke:#5a8f00,color:#fff
style VCF fill:#FFE082,stroke:#FFA000
style GPU fill:#1B2333,stroke:#76B900,color:#76B900Click to view the full pipeline logical diagram
Output¶
The stage produces a VCF file (Variant Call Format) containing:
- Chromosome position of each variant
- Reference allele (what the reference genome has)
- Alternate allele (what the patient has)
- Quality scores (confidence in the call)
- Genotype (heterozygous or homozygous)
This VCF file becomes the input for Stage 2: Evidence RAG, where we determine which variants actually matter.
Why GPU Acceleration Matters¶
Traditional genomics pipelines run on CPU clusters and take 1–2 days per patient. For a hospital processing hundreds of patients, this creates bottlenecks.
GPU acceleration changes the equation:
- Same accuracy — FDA-cleared Parabricks matches CPU results
- 10x faster — Hours instead of days
- Lower cost — One DGX Spark vs. a CPU cluster
- Real-time capability — Results while the patient is still in clinic