Architecture Guide¶
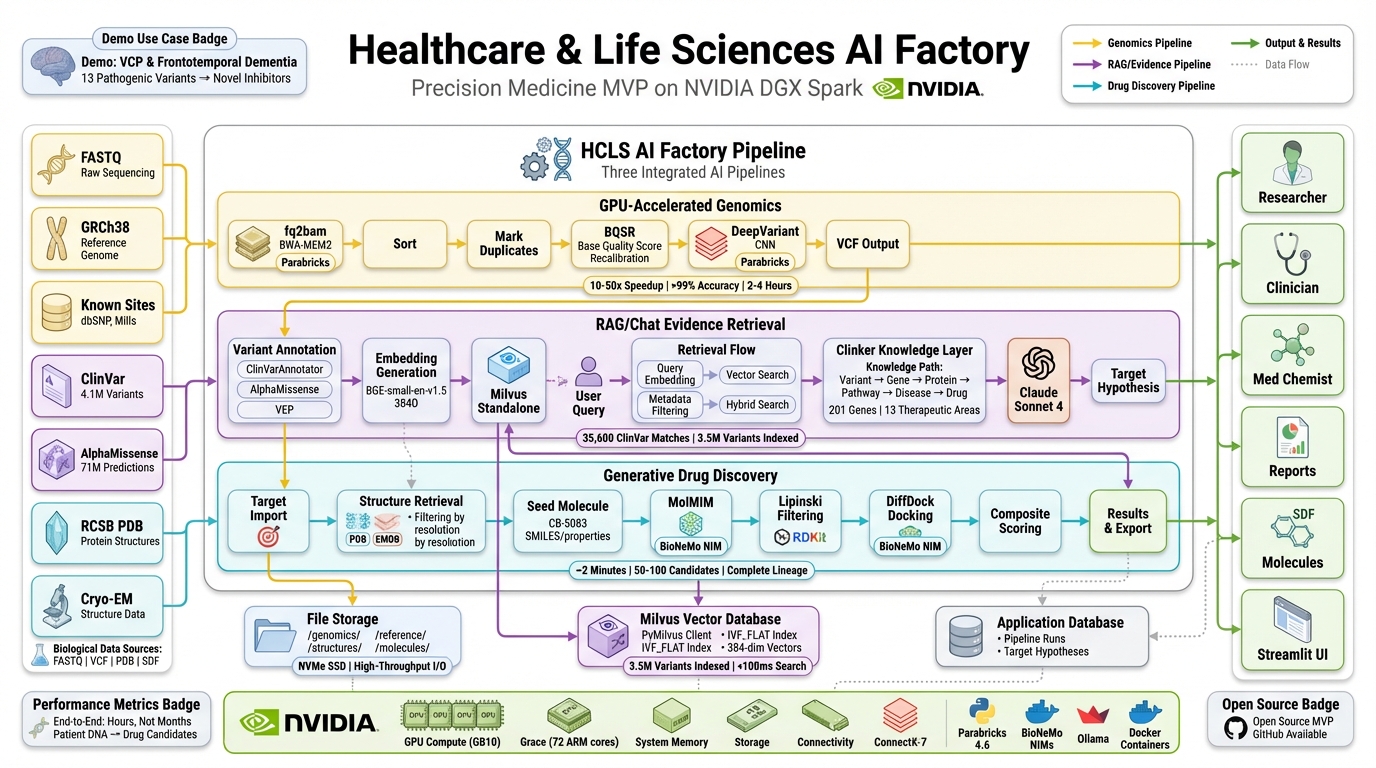
The HCLS AI Factory runs three GPU-accelerated AI pipelines on a single NVIDIA DGX Spark workstation — from raw FASTQ sequencing data to ranked drug candidates in under 5 hours.
Processing Stages¶
Stage 0 — Data Acquisition (one-time): Before the pipeline can run, all required data must be downloaded: HG002 FASTQ sequencing files (~200 GB), the GRCh38 reference genome with BWA-MEM2 index (~11 GB), ClinVar clinical annotations (4.1M variants), and AlphaMissense pathogenicity predictions (71M variants). The setup-data.sh script automates this entire process with checksum verification, parallel downloads, and idempotent resumption. This is a one-time step (~500 GB total).
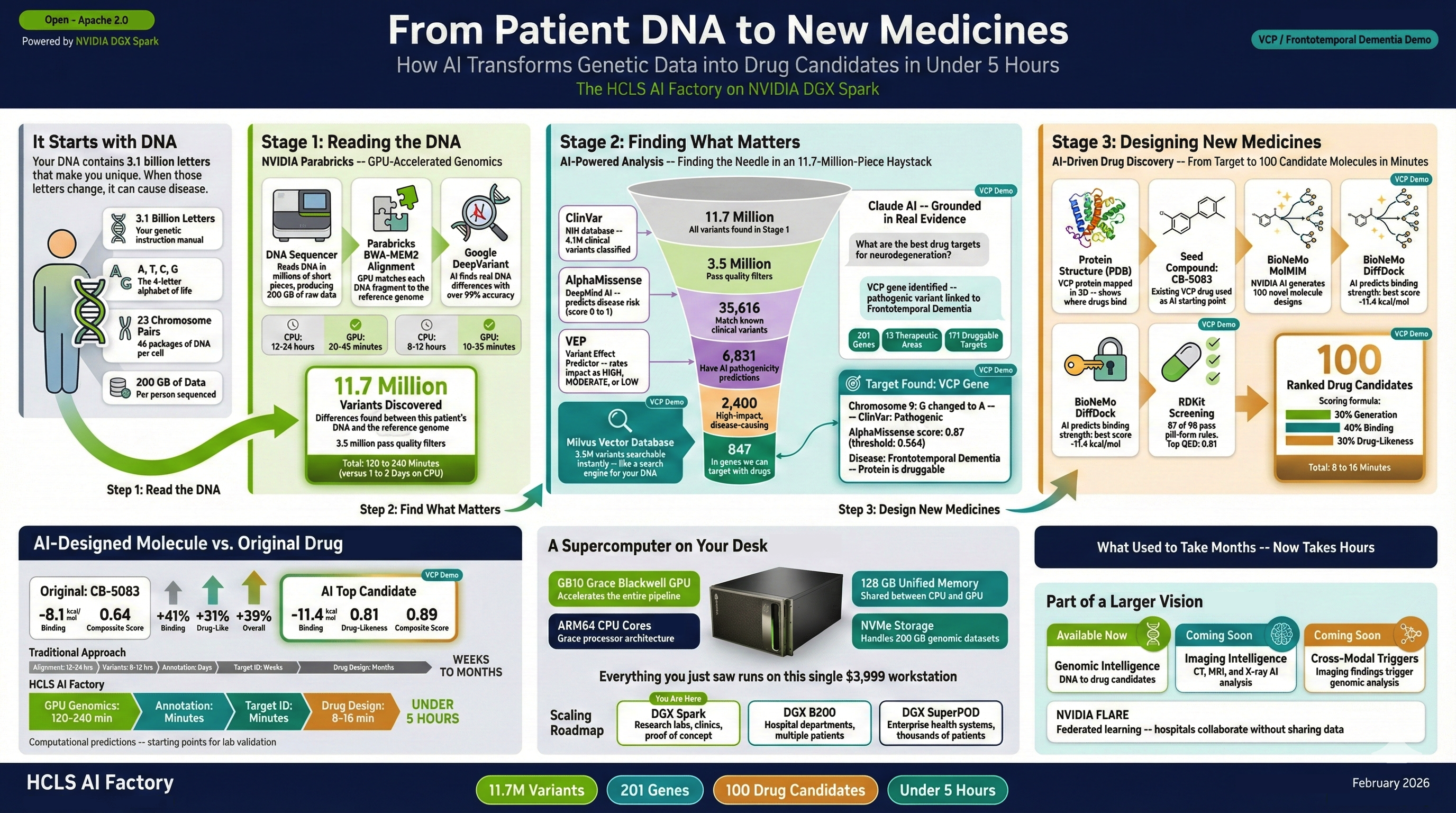
Stage 1 — GPU Genomics (120–240 min): Raw FASTQ sequencing files are aligned to the human reference genome using BWA-MEM2, then variant-called with Google DeepVariant — both accelerated through NVIDIA Parabricks. The output is a clinical-grade VCF containing 11.7 million variants at >99% accuracy.
Stage 2 — Evidence RAG (interactive): Variants are annotated against ClinVar (4.1M clinical records) and AlphaMissense (71M pathogenicity predictions), embedded with BGE-small-en-v1.5, and indexed into a Milvus vector database (3.56M vectors). A conversational RAG interface powered by Anthropic Claude lets researchers query variants in natural language, with every answer grounded in retrieved clinical evidence.
Stage 3 — Drug Discovery (8–16 min): For validated targets, NVIDIA BioNeMo MolMIM generates novel molecular candidates from a seed compound, DiffDock predicts protein-ligand binding poses, and RDKit scores drug-likeness (QED, Lipinski, synthetic accessibility). The result: 100 ranked candidates per target with full structural and pharmacological profiles.
All three stages run on the DGX Spark's GB10 Grace Blackwell Superchip with 128 GB unified memory — connected via NVLink-C2C so the GPU and CPU share the same memory pool without transfer bottlenecks.
Architectural Infographic¶
From Patient DNA to New Medicine Infographic¶
Pipeline Mindmap¶
Hardware Platform¶
| Component | Specification |
|---|---|
| System | NVIDIA DGX Spark |
| GPU | GB10 Grace Blackwell Superchip |
| Memory | 128 GB unified LPDDR5x |
| CPU | ARM64 cores (Grace) |
| Storage | NVMe SSD |
| Price | $4,699 |
Deep Dives¶
- Project Bible — Complete technical reference with scoring formulas, thresholds, and configurations
- White Paper — Architecture overview and design rationale
Clinical Decision Support Disclaimer
The HCLS AI Factory platform and its components are clinical decision support research tools. It is not FDA-cleared and is not intended as a standalone diagnostic device. All recommendations should be reviewed by qualified healthcare professionals. Apache 2.0 License.